11.05.2026
Dark Proteome Unlocked: HLA-Compass AI Finds 17,082 Cancer Targets
11.05.2026

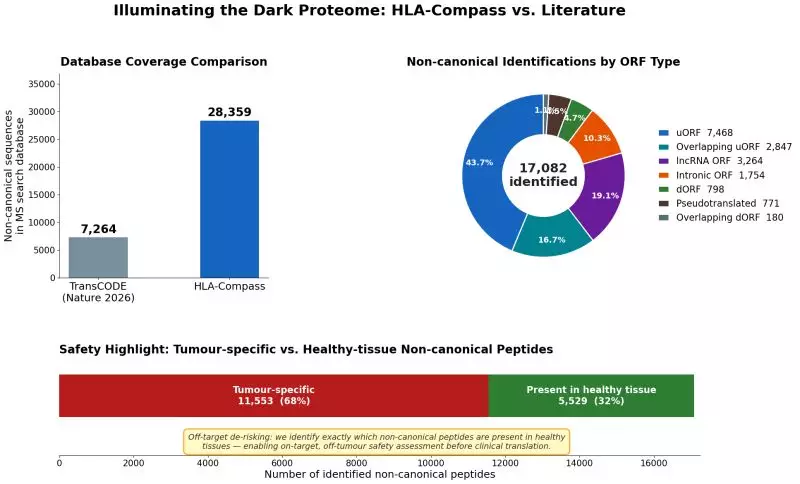
Dark proteome research has just reached a breakthrough milestone. Alithea Bio used HLA-Compass AI to identify 17,082 non-canonical peptides — including 11,553 tumour-specific targets — in just 7 hours. Here is what this means for TCR drugs and cancer vaccines.
The dark proteome refers to the vast set of proteins and peptides encoded by non-canonical regions of the human genome — sequences that traditional reference databases have long considered “non-coding” or biologically irrelevant. These include upstream open reading frames (uORFs), downstream ORFs (dORFs), long non-coding RNA-derived ORFs (lncRNA-ORFs), and other non-canonical open reading frames (ncORFs) that fall outside the approximately 19,500 canonical protein-coding genes.
For decades, these dark proteome regions were largely ignored in drug discovery. But a growing body of evidence — and now a landmark study published in Nature — confirms that the dark proteome is not empty. It is, in fact, teeming with biologically active molecules that could reshape cancer immunotherapy.
In May 2026, the TransCODE Consortium published “Expanding the Human Proteome with Microproteins and Peptideins” in Nature — one of the most significant dark proteome papers in recent years.
The study, led by researchers including Eric Deutsch, Steven Carr, Jenn Abelin, Pouya Faridi, Nicola Ternette, John Prensner, and Sebastiaan van Heesch, analysed over 7,264 non-canonical open reading frames (ncORFs) across 95,520 proteomics experiments. Their core finding: approximately 25% of those ncORFs produce detectable peptides — confirming translation of sequences that have never appeared in standard protein databases.
The team identified around 1,785 previously unrecognised microproteins, introducing the new conceptual category of peptideins — microproteins under 50 amino acids with unusual properties, some of which are essential for cell survival and implicated in cancer biology. The human proteome, the paper concludes, is nearly 10% larger than previously recognised.
As co-lead Sebastiaan van Heesch noted, thousands of overlooked genetic sequences contribute to a new class of protein-like molecules that had been missed until now. All data were shared open-source to accelerate global research.
External reference: Nature — “Expanding the human proteome with microproteins and peptideins”
After reading the Nature paper, Alithea Bio’s scientists — Alisa Fuchs, PhD and Armanas Povilionis — immediately asked a more specific question: What happens when you search the world’s largest HLA peptide database against the full dark proteome catalogue?
Rather than relying on the 7,264-sequence initial catalogue from the Nature study, the team went further. They used a revised, expanded catalogue from Chothani 2026, comprising 28,359 non-canonical ORF sequences — covering lncRNAs, uORFs, dORFs, downstream overlapping ORFs, and more.
Using HLA-Compass AI’s fully automated reanalysis pipeline, the team searched thousands of real-world immunopeptidomics samples from the HLA-Compass database — one of the largest collections of healthy tissue, cancer cell line, and tumour tissue HLA peptidomics data in the world.
The results were ready in just 7 hours.
The dark proteome numbers are transformative:
| Metric | Result |
|---|---|
| Non-canonical peptides identified | 17,082 unique peptides |
| Tumour-specific peptides | 11,553 (not found in any healthy tissue) |
| ORF types covered | lncRNAs, uORFs, dORFs, and more |
| Catalogue used | Chothani 2026 — 28,359 sequences |
| Time to results | 7 hours |
| Platform | HLA-Compass AI automated reanalysis pipeline |
The most striking finding: 11,553 of those dark proteome peptides are tumour-specific — meaning they appear in cancer tissue but are absent from any healthy organ in the database. This represents a largely unexplored reservoir for TCR therapy development, cancer vaccine design, and neoantigen discovery.
The discovery of thousands of tumour-specific, non-canonical dark proteome peptides has direct implications for multiple drug development programmes:
T-cell receptor (TCR) therapies require high-confidence tumour-specific targets that are absent from healthy tissue. Off-target expression in vital organs is a primary safety concern. Dark proteome cancer targets — peptides derived from lncRNAs, uORFs, and dORFs that appear only in tumour tissue — represent a new class of candidates with a naturally favourable safety profile. HLA-Compass AI’s absolute quantification data allows researchers to verify both tumour presence and healthy tissue absence at scale, before committing to expensive experimental validation.
Personalised cancer vaccines depend on identifying neoantigens and tumour-associated antigens that the immune system can be trained to recognise. Non-canonical dark proteome peptides are particularly attractive vaccine targets because they are derived from sequences with no normal protein counterpart — reducing the risk of central tolerance and autoimmunity. The 11,553 tumour-specific peptides identified by Alithea Bio’s pipeline expand the universe of vaccine candidates dramatically.
Traditional neoantigen pipelines focus on somatic mutations in canonical coding sequences. The dark proteome opens a parallel pathway: non-canonical sequences that are transcribed and translated specifically in cancer cells, independently of mutation status. This matters especially for tumour types with low mutational burden, where canonical neoantigen identification is difficult.
Neoantigens | NeoZOOM | HLA-Compass | Off-Target Toxicity Prediction
The dark proteome comprises several categories of non-canonical open reading frames:
lncRNA-ORFs — Open reading frames embedded within long non-coding RNAs. These transcripts were historically assumed to have regulatory roles without producing proteins. Evidence now shows many are translated, producing small proteins with functional relevance in cancer biology.
uORFs (Upstream Open Reading Frames) — Short ORFs located in the 5′ untranslated region upstream of a canonical coding sequence. They can act as translational regulators and, when expressed aberrantly in cancer, produce peptides that may be presented on HLA molecules.
dORFs (Downstream Open Reading Frames) — ORFs located downstream of or overlapping with the main coding sequence. Like uORFs, they are translated in specific cellular contexts and may generate tumour-associated peptides invisible to standard proteomics approaches.
Together, these non-canonical sequences constitute the dark proteome — a parallel protein landscape that has been systematically overlooked until now.
HLA-Compass AI is Alithea Bio’s SaaS platform — the world’s largest quantitative HLA peptide database, combined with an AI-powered analysis engine purpose-built for dark proteome discovery. Its key capabilities include:
The pipeline’s ability to complete a full dark proteome reanalysis against 28,359 sequences in just 7 hours is a direct result of this AI-powered infrastructure.
One of the most powerful outcomes of this work is that HLA-Compass AI’s pipeline can be applied to any dark proteome sequence database — not only Alithea’s own catalogue.
If your team has developed a proprietary catalogue of non-canonical ORF sequences, you no longer need to wait months for experimental mass spectrometry validation. Submit your catalogue to Alithea Bio’s pipeline and receive results against thousands of real-world healthy and cancer samples — delivered the next day.
To learn more, reach out via the contact page, or meet the team at PEGS Boston and the TCR T Cell Congress.
The dark proteome is the collection of proteins and peptides encoded by non-canonical regions of the genome — including lncRNAs, uORFs, and dORFs — absent from standard protein reference databases. A 2026 Nature study confirmed that approximately 25% of 7,264 such regions produce detectable proteins, expanding the known human proteome by nearly 10%.
Peptideins are a newly defined class of dark proteome microproteins — under 50 amino acids — encoded by non-canonical open reading frames. Formally defined in the 2026 TransCODE Consortium paper in Nature, many are now being investigated as drug targets and disease biomarkers.
Alithea Bio identified 11,553 tumour-specific dark proteome peptides — present in cancer tissue but absent from all healthy tissue samples in the HLA-Compass database — out of 17,082 non-canonical peptides detected in total.
Using HLA-Compass AI’s automated pipeline, Alithea Bio can search any dark proteome sequence database against thousands of real-world immunopeptidomics samples and deliver results the next day.
Yes. Alithea Bio has optimised its pipeline to accept any protein sequence database. Contact the team via alithea-bio.com/contact to discuss your project.
HLA-Compass AI is Alithea Bio’s SaaS platform for navigating the human immunopeptidome and dark proteome. It combines the world’s largest quantitative HLA peptide database with AI-powered tools for peptide-HLA binding prediction, off-target toxicity assessment, and automated database reanalysis.
Meet the team at PEGS Boston and the TCR T Cell Congress — or get in touch to discuss how HLA-Compass AI can accelerate your dark proteome programme.
Related reading: HLA Presented Peptides | Neoantigens | Immunopeptidomics Technology | Proteogenomics